Solubility prediction - Chemaxon's Solubility Predictor
Aqueous solubility for organic compounds is one of the most important physico-chemical properties in modern drug discovery and has long been among our plans. Being a property that is hard to model, providing a tool for prediction, in a fast and accurate way, has not been an easy task and so we are pleased to announce our solution in the latest 6.3 release.
Chemaxon's Solubility Predictor is able to predict aqueous intrinsic solubility and pH-solubility profile for molecules. An example below shows the prediction results for the acebutolol molecule.
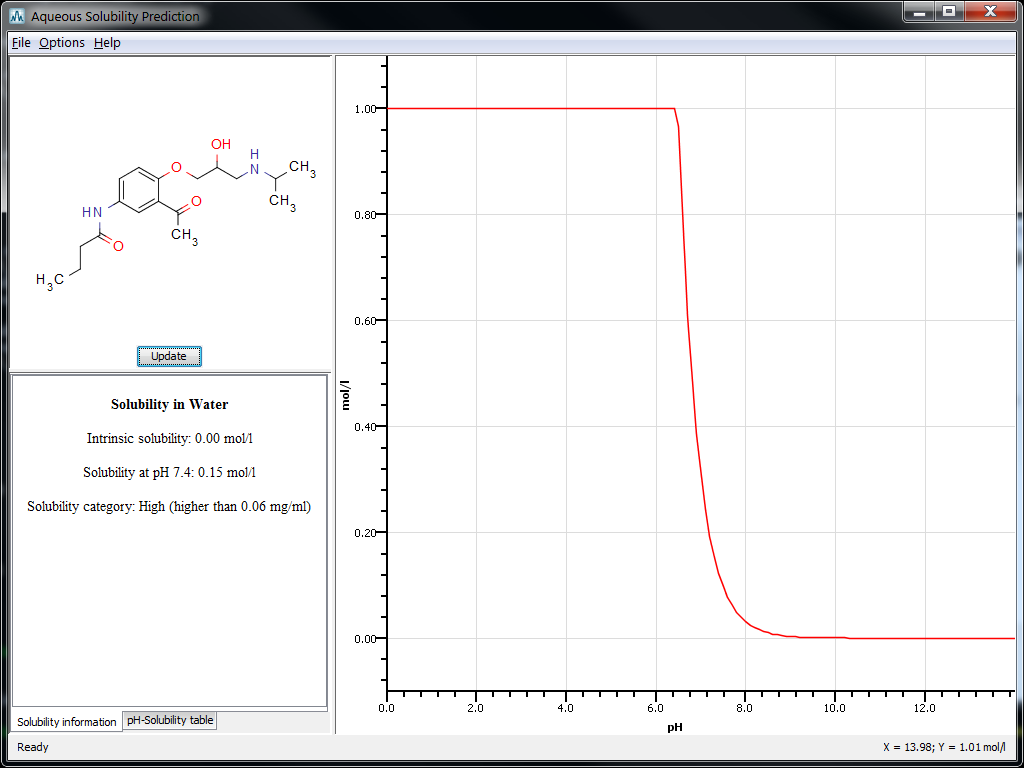
The performance of the intrinsic solubility predictor was measured using the R2 value for the training set (0.91). This is similar to the value in the references on which the implementation was based, showing good average performance of the predictor.
You can use Solubility Predictor as a plugin via the MarvinSketch GUI and as a calculation in cxcalc. It is also available in Instant JChem, JChem for Office Suite and as a Chemical Terms function. Using solubility functions via Chemical Terms from Instant JChem or JChem for Office is a nice way to perform filtering based on solubility that can be used in ADMET predictions.
For example if you want to filter a library of compounds based on solubility at pH = 7.4, you should use the following filter that is evaluated for each molecule in the library, and updated if a library is edited:
evaluate -e "logS('7.4') > 2.5" library.sdf
An other useful filter can be made based on how well soluble the compound is :
evaluate -e "logSCategory() == 'high'" library.sdf
If you would like to know more about Chemaxon's Solubility Predictor, visit the product page or find details in the user guide.
Related content
Certainty Discovery. Frankfurt, November 4-5, 2025
Certainty Discovery 2025 review by Wendy A. Warr - Vibes, topics and key takeaways.
The Rise of Biologics Discovery: From Small Molecules to Sequences
Biologics in drug discovery is becoming more important by the year. See where biologics discovery...
Tautomer Generation Methods – Case Study on Reliability
Tautomer data handling in a reliable manner - case study based on thorough testing to find the most...