Local and Global Models For Predicting Properties of Small Molecules
To be most successful, which type of model should be built?
The obvious answer is all models that are prospectively predictive. Also, it is very valuable if the models are interpretable, to facilitate design of new compounds with improved properties.
Another important parameter is the size of the applicability domain. A large domain implies that the input compounds can be highly diverse, but still recognized and correctly predicted by the model, thus allowing for extrapolating into a more promising chemical space.
How would a medicinal or computational chemist know which type of model to build, to be most successful?
The choice between local and global models
Usually, there is a balance between applicability domain and predictive power, where a larger domain (a global model) often comes at the expense of less accurate predictions, which are easier to achieve with a more local model (Figure 1).
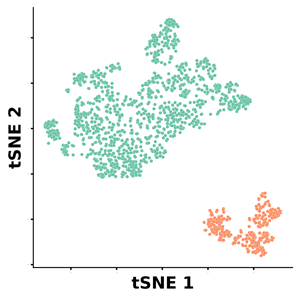
Figure 1. The size of the applicability domain of a global and a local model is relative and depends on the training set used. The key aspect is not the domain size but rather whether the model is valid for the molecules of interest.
Workflow for creating predictive models and the benefits of continuously adding new data
Publishing models for medicinal chemists
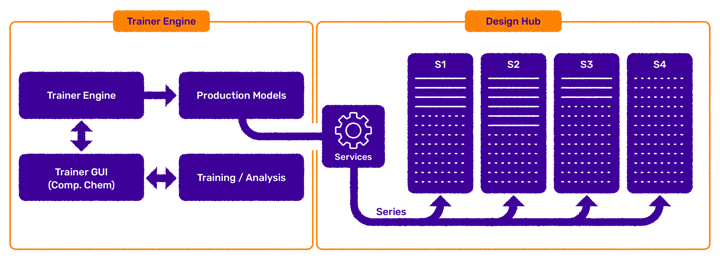
Figure 2. Integration overview. Validated, production-grade predictions can be made available as a Design Hub plugin to foster selecting the most viable idea molecules and novel designs (Fig. 2.).
Comparing local and global models
Simulation of (Fig. 3.) local and global model building with re-training and comparison.
- A ‘random test’ set (203 cases) is selected from the initial dataset (2029 cases). [1]
- Step two selects a scaffold cluster (146 cases) and provides the rest of the compounds (1680 cases) as a ‘Global’ set. The largest Tanimoto similarity between the scaffold set and the global set is 0.784.
- The scaffold set is divided into a training set (102 cases) and in a consecutive step into an ‘update’ and a ‘final test’ set (22 cases each).
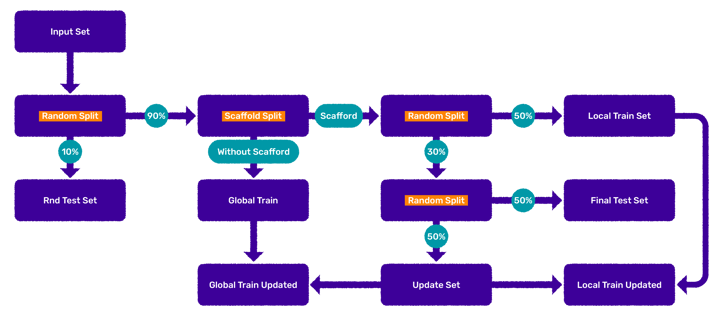
Figure 3. Data preparation workflow
Observations
- Random Forest binary classification model was built using 19 selected descriptors for each set.
- Global model performed better on randomly selected external test set (Fig. 4.) compared to the Local model.
- Although the Local model is trained on 16x less data, it outperformed the Global model on the analogues of the scaffold set (Fig. 5.).
- Re-training both models with additional scaffold analogues improves performance on second test set of scaffold derivatives (Fig. 6.)
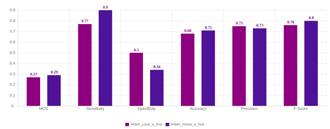
Figure 4. Local and global model performance on external data

Figure 5. Performance tested on scaffold analogues
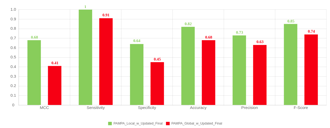
Figure 6. Re-trained model performance tested on second round of scaffold analogues
—
[1] NCATS Parallel Artificial Membrane Permeability Assay (PAMPA) (1508612)
Related content
Certainty Discovery. Frankfurt, November 4-5, 2025
Certainty Discovery 2025 review by Wendy A. Warr - Vibes, topics and key takeaways.
The Rise of Biologics Discovery: From Small Molecules to Sequences
Biologics in drug discovery is becoming more important by the year. See where biologics discovery...
Tautomer Generation Methods – Case Study on Reliability
Tautomer data handling in a reliable manner - case study based on thorough testing to find the most...