Key Properties in Drug Design | Predicting Lipophilicity, pKa and Solubility
PhysChem properties are key descriptors in drug design
Lipophilicity, pKa and solubility are key descriptors in drug design. Their importance to both pharmacokinetic exposure (ADME) and pharmacodynamic response (effect on target and off-targets) have been thoroughly studied and described in the literature (Figure 1).
Most drug discovery companies are measuring these properties in a high-throughput manner and drug designers are frequently predicting them prior to prioritizing compounds for synthesis.
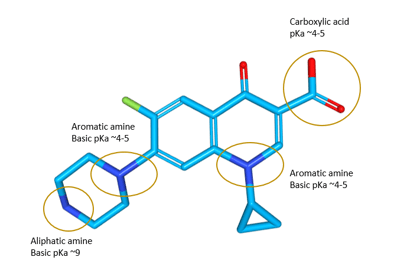 Figure 1a. The pKa of a drug directly influences its ionization state which is correlated to lipophilicity, solubility, affinity to proteins (on and off-targets) and permeability across membranes. Also crucial to ionization state is pH, which is varying from the highly acidic environment of the guts (pH ~2) across the weakly acidic lysosomes (pH ~5) to the physiologically normal intracellular pH in most cells (pH ~7.5). The majority of drugs are weak acids and/or bases. [2]
Figure 1a. The pKa of a drug directly influences its ionization state which is correlated to lipophilicity, solubility, affinity to proteins (on and off-targets) and permeability across membranes. Also crucial to ionization state is pH, which is varying from the highly acidic environment of the guts (pH ~2) across the weakly acidic lysosomes (pH ~5) to the physiologically normal intracellular pH in most cells (pH ~7.5). The majority of drugs are weak acids and/or bases. [2] Figure 1b. Solubility in water is essential for pharmacokinetic exposure, since the drug needs to dissolve in the aqueous body fluids to reach the target tissue. Solubility is also crucial for maintaining an appropriate concentration in any in vitro experiments, thus avoiding artefacts from for instance aggregation.
Figure 1b. Solubility in water is essential for pharmacokinetic exposure, since the drug needs to dissolve in the aqueous body fluids to reach the target tissue. Solubility is also crucial for maintaining an appropriate concentration in any in vitro experiments, thus avoiding artefacts from for instance aggregation.
Figure 1c. Lipophilicity is usually measured by the octanol/water partitioning coefficient. Highly lipophilic compounds have an increased risk of being poorly soluble, quickly metabolized and promiscuous binders, whereas more polar compounds often are less permeable. Thus, lipophilicity is an important parameter for all ADMET properties of a drug [4].
Predictive Models for pKa, Solubility, and Lipophilicity: examples and model performance
Lipophilicity, pKa and solubility are well suited parameters to be predicted by machine-learning approaches, owing to the large amount of high quality data available. Chemaxon have developed high performance predictive models for all three parameters (Figure 2, 3, 4 and 5).
Our “Calculated properties” tool include not only predictions of the three key physchem parameters (cLogP, solubility and pKa) but also structure-based calculations of other commonly used molecular descriptors such as molecular weight, polar surface area, fraction sp3 carbons, number of hydrogen-bond acceptors and donors. Calculated properties for designed compounds can be compared to a reference.
| Calculated properties | |||
| Current | Pinned | ||
| ▲ | Mass | 478.52 | 474.48 |
| ▲ | cLogP | 6.63 | 3.38 |
| ▼ | TPSA (Ų) | 51.36 | 87.46 |
| ▼ | pKa (str. acidic) | 13.48 | 13.57 |
| ▲ | pKa (str. basic) | 8.8 | 2.07 |
| ▼ | FSP3 | 0.26 | 0.38 |
| ▼ | Solubility (mM) | 0 | 0.01 |
| ▼ | H-bond acceptors | 3 | 4 |
| ▼ | H-bond donors | 1 | 2 |
Figure 2. Calculation of cLogP, solubility, pKa and other various molecular descriptors
Performance of the logP model
The Chemaxon calculators yielded in the lowest RMSE, MAE and the highest R2 compared to other participants, ranking it as the most accurate predictor. On this 11 novel compound dataset, only one structure had higher than 0.5 unit error. This challenging entry had lower than 1 log unit deviation comparing measured and calculated data. Learn more from our whitepaper.
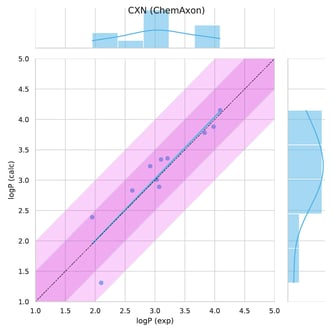 Figure 3. Performance of the logP model is shown on SAMPL6 blind data.
Figure 3. Performance of the logP model is shown on SAMPL6 blind data.
Accuracy of thermodynamic solubility prediction
The accuracy of thermodynamic solubility prediction in water was assessed on a dataset with 6886 compounds. In 46% of the cases the predicted solubility was found to be within 0.5 logS unit and in 75% of the cases the error was less than 1 logS unit. This concludes to 1.04 RMSE and 0.86 Pearson correlation coefficient. For details and references please read our corresponding whitepaper.
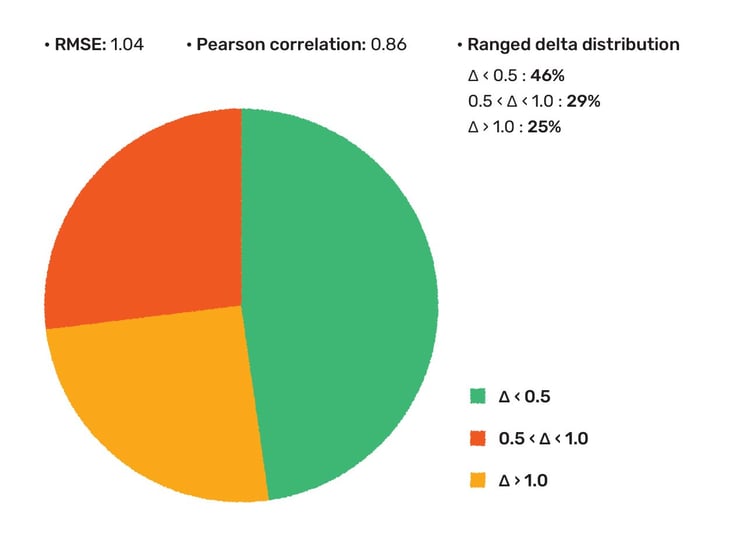 Figure 4. The accuracy of thermodynamic solubility prediction evaluated on a drug discovery set
Figure 4. The accuracy of thermodynamic solubility prediction evaluated on a drug discovery set
SAMPL7 blind challenge results
According to the published results on the blind challenge of pKa prediction, Chemaxon algorithm had the lowest RMSE on the SAMPL7 challenge. The original authors concluded that: “We tested Chemaxon’s Chemicalize toolkit as an empirical reference method to make macroscopic pKa predictions and it performed better than other methods.“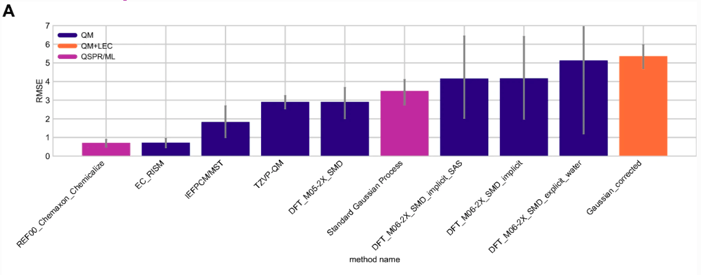
Figure 5. Evaluation of molecular predictions from the SAMPL7 blind challenge
[1] Bunally Using Physicochemical Measurements to Influence Better Compound Design. SLAS 2019;24(8):791-801.
[2] Manallack DT. The pK(a) Distribution of Drugs: Application to Drug Discovery. Perspect Medicin Chem. 2007;1:25-38.
[3] Bergström et al, Drug solubility in water-based systems International Journal of Pharmaceutics 540 (2018) 185–193
[4] Waring, Lipophilicity in drug discovery Expert Opin Drug Discov. 2010 Mar;5(3):235-48.