Wendy Warr Report 2022
CHEMAXON EUROPEAN USER MEETING, BUDAPEST, MAY 29-31, 2022
A report by Wendy Warr (https://www.warr.com/)
Introduction
With pandemic restrictions lifted, Chemaxon users yet again convened in person at the Akvárium Club, Budapest. The event began with a social gathering at the club on the Sunday night. The gala dinner on the Monday night was at a surprise venue out of town, in an unusual industrial setting, with fire-eating dancers and fascinating metalwork sculptures. The after-dinner music was deafening as usual but there was an early bus home for the faint-hearted.
Chemaxon and Its Portfolio
Moving forward together with Chemaxon
Richard Jones, CEO of Chemaxon reminded us that Ferenc Csizmadia, founder of Chemaxon, said the company was founded on innovation with approachability and eagerness to help, combined with honesty, openness, and transparency. Marvin JS was all about accessibility for chemists: the market-leading structure editor had to be installed on your computer, but Marvin JS could be run on your browser. The portfolio, said Richard, was expanded to other fundamental solutions used every day and Chemaxon has come to be recognized as the best in drawing, search, calculators, compliance, and standardization.
Innovations were web-readiness (even in the 1990s), a full-coverage cheminformatics toolkit, accurate and fast structure-based predictions and structure searching, Markush search, and an easy-to-use API. Many workflows and differing business logic options are supported. The accessible, innovative approach quickly led to Chemaxon being adopted everywhere chemistry is being performed. The software has more than 1 million users, not only in pharma and biotechnology industries but also in agrochemicals, flavors, publishing, and academia. The company employs 220 people (rising to 300 in 2023) in six offices. It has over 70 partners.
Chemaxon is determined to stay ahead, fiercely independent and stable at the cost of external investment. It will focus on game-changing ideas, leading to strong growth but never growing too big for approachability and customer focus. It will continue to focus on close collaboration, following the examples of Instant JChem (developed with Novartis), Compound Registration (with GSK), and Marvin Live, developed with Vertex and leading to Design Hub (with Boehringer Ingelheim).
Chemaxon moved from point solutions to integrated solutions for chemistry and biology and is now covering larger workflows with end-to-end cloud solutions. It has a best-in-class, single research platform in the cloud for end-to-end, early-phase drug discovery. On top of everything are custom developments, technological partnerships, training, first-class support, and data services, not just in industry but also in chemistry education.
The strategy for 2025 is to cultivate a progressive company culture where everyone contributes to company growth, builds a best-in-class product and service experience, solves customers’ needs on the cloud, and serves the early phase R&D of biopharmaceuticals. Chemaxon’s values are customer excellency, integrity, long-term thinking, and collaboration.
New challenges, new direction: portfolio overview 2022
“We are your trusted partner to build a better future through innovative best-in-class software for chemistry and biology.”
Efi Hoffman, Head of Research Platform at Chemaxon, gave an overview of changes in the pharmaceutical industry which affect daily R&D work and needs, leading to opportunities for Chemaxon to develop products further and to devise newer and better ways to serve their customers.
A major trend relates to data. In 2006, Clive Humby coined the phrase “data is [sic] the new oil” and Michael Palmer expanded on this by saying, that like oil, data are “valuable, but if unrefined, cannot really be used”. Most of the problems (different formats, conversion, management of migration) concern legacy data, but even new data should be collected or standardized in a unified format. The Findable, Accessible, Interoperable, and Reusable (FAIR) movement started more than six years ago, but we are still not able to solve these problems. Security is a big issue with data. Another is handling, storing, and finding relevant information in sources which contain structured and unstructured data together. Data lakes are very popular nowadays, but there is still a lot of room for improvement here.
Trends in outsourcing, CROs, and collaborations also present opportunities for Chemaxon. The CRO market is growing extremely fast: it had already reached $50 billion by 2021 and a compound annual growth rate of 8.5% is expected from 2021 to 2028. Merger and acquisition business started to grow again in 2021; research groups in new or extended organizations need to share information and work together. There are also 498 consortia in pharma facing the challenge of sharing data.
Other game-changers in pharma are new domains. Biological drugs are making up a growing share of FDA approvals. Drug delivery technologies have had to change.1 Researchers need different parameters to optimize, synthesize, and formulate the drugs, and software requirements are different from those for small molecules.
Software users are also changing their habits. They access their emails, favorite music, and books from different devices; they use the touch screen of their mobile; they give sound commands to their phone or Alexa. They will not accept anything less from their laboratory software. The Pistoia Alliance initiative “User Experience for Life Sciences” (UXLS) helps the life science R&D community to understand UX skill sets.
Finally, Efi summarized the unsolved challenge of the application of artificial intelligence in pharma. Sixteen billion dollars have been spent on AI development in the pharma industry from 2010 to 2021, yet there has been no huge increase in the number of drug candidates in the clinical phase. A recent review2 suggests that Al has had more relevance in early-phase drug discovery projects, and less in phases I, II, and III. Andreas Bender concludes that when it comes to computational modeling of data, a method cannot save an unsuitable representation, which cannot remedy irrelevant data for an ill thought-through question. We still need to work on effective AI software.
Next, Csaba Peltz briefly discussed how Chemaxon can address the challenges presented by Efi. As regards data issues, and changing user habits, needs, and expectations, Chemaxon is now moving into the education market, starting with Zosimos. Csaba repeated Richard Jones’ comments about the platform direction. It is called a platform because it is modular; it is consistent in UI and UX, component interactions, security, design patterns and principles; it can be extended with new components (even by customers); and it is used to create products by combination and configuration of components.
Point solutions in 2022 addressing data, AI, and collaboration challenges include Calculators, molecular descriptors, Trainer Engine, Reactor, and Compliance Checker and cHemTS. Addressing data and UX challenges are MarvinSketch, Marvin JS, Marvin Pro, Standardizer, Structure Checker, Markush Editor, Markush Enumeration, Markush Search, Naming, Document to Structure, and ChemCurator. Data are handled by JChem Engines and Hosted Catalog Search. Integrated solutions that address multiple challenges are Design Hub, Instant JChem and Plexus Connect, JChem for Office, DataLink, Compound Registration, Biomolecule Toolkit, and the Small Molecule Synthetic Chemistry (ELN) solution. Csaba introduced the product managers, and speakers from the user community, related to those products specifically covered in the user meeting program.
Laboratory Workflow
From idea to insights: data capture across a laboratory workflow
Roland Knispel, Product Manager, Chemaxon
By adding an ELN, Chemaxon now covers the whole of the design, make, test, analyze (DMTA) cycle (Figure 1). Chemists can sketch a reaction step in Marvin JS and get a table which helps in setting up the reaction. They specify the intended stoichiometry of the reaction by adding the equivalents. Preconfigured formulas take care of calculating the ingredients. Theoretical moles, masses, and volumes are calculated for the reactants, considering purity. Reactants can be selected from the compound inventory. Documentation is simplified with dynamic text building blocks.
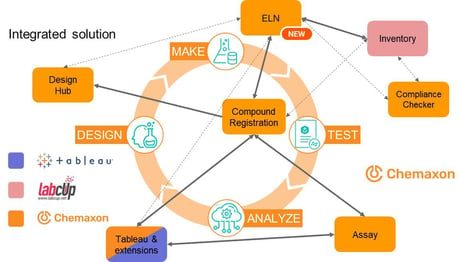
Customizable forms to request analytical measurements are available. Requests are tracked and the status of a request is notified. Analytical results can be imported, and files attached. The product can then be registered in Compound Registration. Chemists can contact a CRO and request synthesis of a library. When the synthesis is completed, the CRO partner sends the material, along with an SDfile or text file containing the chemical data. During import into Compound Registration, data fields are mapped and the regular standardization and structure quality checks follow. Biological assays are now requested, and the request and results are handled by Chemaxon Assay. ADME requests can be sent to a CRO, and results can be automatically uploaded from cloud drives.
Chemaxon has added extensions to Tableau for data analysis. For example, SAR tables can be constructed and viewed. Here, compounds with a balanced profile can be identified and similar compounds can be found (with the structure search extension). The card view extension allows users to compare structures by showing them side by side. On the scatter plot dashboard, chemists can select a portion of the data from a table and view the plot above the table (useful for monitoring outliers, for example). There is also a correlation analysis dashboard. Finally, users can see a summary of the data for a given entity in the compound résumé.
This is a research informatics cloud platform centered on medicinal chemistry needs, now extended by an ELN for tracking chemical syntheses, with a strong emphasis on workflow-driven UX design and development. Chemaxon is looking for early adopter SMEs to try it out.
LabCup inventory management software
Gábor Radics, CEO, LabCup
LabCup is a modular, integrated, inventory management system for research samples, consumables, and lab equipment. It is scalable to multiple sites, countries, and hosts. There can be more than 30,000 users per host. LabCup’s key principles are automation; using and processing raw data; using the newest technology; prioritizing the user experience; and development with the customer. The system supports the entire chemical life cycle: purchasing, loading, registration, storage, compliance, and waste disposal. Gábor outlined features for scalability, saving of storage space, automation, and the use of mobile devices.
Marvin JS is integrated in the federated search functionality of LabCup and the Chemaxon ELN (see above) is also integrated. Current stock information, compound registration, physicochemical properties, safety information, and analytical equipment booking are thus featured. The user saves time through this integration because data such as synonyms, chemical structures, safety data sheets and safety information are automatically collected; scanning can be used in stock taking; pictures can be imported; and compliance reports can be generated. The data can be processed for segregating chemicals by storage location and estimating storage risk. A user company’s environment, health, and safety (EHS) office can set maximum thresholds in labs and LabCup will highlight breached limits by color. Chemical data are pulled directly into the risk assessment module from the chemicals datasheet. The safety officer receives risk assessments for approval if an item is regulated.
Chemaxon software is built into the chemical inventory, equipment booking and asset management, EHS, and risk assessment modules of LabCup. Other LabCup modules include digital floor plans with live emergency and hazard information, hazardous waste disposal, a purchasing platform, and API integration. Chemaxon customers can benefit from significant cost savings in the full chemical life cycle. Storage costs are saved because chemicals can be shared and are not built up. Total costs of ownership are reduced, instruments can be shared, and less training is needed for integrated software. A large-scale implementation of LabCup has been reported recently.3
Marvin Pro: news and plans about chemical drawing
Bertalan (Berci) Kovács-Garai, Product Manager, Chemaxon
The vision for Marvin Pro is that of a universal chemical editor component, capable of drawing big synthesis schemes, and biomolecules, in publication quality, which can be easily extended with other modules, and can be integrated with Chemaxon’s products and those of other vendors. MarvinSketch is for use on the desktop alone while Marvin JS is a lightweight web application. Marvin Pro is meant to serve both markets. The design principles are impeccable visual quality, fast drawing, and chemical smartness. Berci demonstrated how fast drawing is achieved by use of the keyboard and precision alignment. He also illustrated smartness with display of physicochemical properties on the fly, 2D cleaning, stereo calculation, aromatization, easy name input, and handling of common abbreviations for substructures such as functional groups.
Content can be copied and pasted in various formats (V2000 and V3000 molfile, MRV, SMILES, CDX, SKC, and InChI), is compatible with other drawing tools, and can be exported in image or chemical file formats. Chemaxon Object Notation (CXON) is a new web-native format, described in more detail later in the meeting. Marvin Pro is an on-premise, hosted web application, with a single installer, where CXON links the Marvin Pro client-side, typescript library with web services on the server.
Features to be added next include atom properties (isotopes, valence, and atom mapping); enumeration features (R-groups, positional variation bonds, and repeating units); search features (query atoms, bonds, and properties); Lewis structures; reaction mechanisms; and stereochemical notation. Further into the future, there will be a customizable user interface, conformance with WCAG 2.1, and fast drawing improved with full keyboard support and drawing large mechanisms. Visuals will be improved further with color schemes and journal presets; there will be support for biopolymers; Office will be integrated; other platforms (Docker, a hosted solution (SaaS), and Marvin Pro on the desktop) will be supported; and structure checking and fixing will be added. Marvin Pro will be integrated in the ELN and Design Hub.
The SaaS version of Marvin Pro should be released at the end of 2022 and the desktop version by Q3 of 2023. The plan is to reach feature parity with Marvin JS in Q1 of 2024 and feature parity with MarvinSketch in Q1 of 2025. Berci closed with a demonstration of publishing your results with Marvin Pro using as an example (Figure 2) a scheme published by the American Chemical Society.4
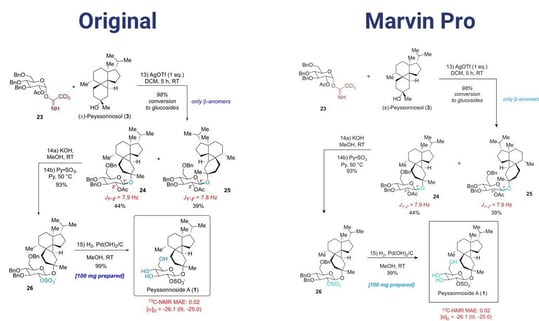
Designing New Molecules
Design Hub at Boehringer Ingelheim
Ferenc Köntés, Principal Scientist and Research Project Leader, Boehringer Ingelheim
The central question for medicinal chemistry design teams is which compound to make next. This question is difficult to answer because drug discovery is complex: druglike chemical space is very large and exploring it requires multiparameter optimization and large-scale data analysis. This was the motivation for Boehringer Ingelheim (BI) to codevelop Design Hub with Chemaxon in 2018-2019. Since then, all BI design teams have adopted Design Hub as their central tool. It allows project overviews, data integration and predictions, and hypothesis-driven drug design in order to prioritize compounds for synthesis. Functions include predictive models, links to external and internal data sources, data-driven recommendations, 3D modeling and docking, and safety alerts.
A survey has shown that it is most used for designing new compounds and sharing designs with team members. It is also used, but to a lesser extent, for reviewing and prioritizing new ideas in design meetings, and editing or contributing to a design set or hypothesis created by other people. It is least used for tracking the status of compounds from design to registration and for synthesis planning and communication with the lab.
Design Hub at BI supports diverse workflows. For example, Project A has no strict rules for when to create a new design set or hypothesis. In preparation for a team meeting, SAR observations and design ideas are captured and there are multiple contributors per design set or hypothesis. During the meeting, new compounds “ready for review” are discussed and selected ones are sent to “for synthesis” and prioritized. After the meeting, one designer transfers compounds to PowerPoint for synthesis planning to be discussed separately. For project A, support for live sessions is critical. There is a backlog of compounds ready for review so an archiving system would be highly desirable. Many design sets and hypotheses are generated, so the ability to organize and keep an overview is critical. BI needs better ways to integrate synthesis planning and lab scientists into daily workflows to avoid time-consuming and error-prone transfer steps.
Project B has a limited number of hypotheses based on structural classes and key recurring topics (e.g., X-ray structures). Design sets are idea sets, not synthesis sets. There is heavy use of comments to share links to outside information (e.g., building blocks and calculation results). In preparation for team meetings, SAR observations and design ideas are saved. During team meetings, new compounds “ready for review” are discussed and selected ones are sent to “for synthesis” and prioritized. Additional ideas from brainstorming sessions are captured. After the team meeting, the lab leader saves synthesis plans in the design set editor and assigns them to lab scientists. Project B has taught BI that limiting the number of hypotheses helps maintain a project overview and including synthesis plans provides high transparency for the whole team. The current editor has only limited functionality for synthesis planning.
The combichem workflow considers one design set as one combinatorial library. A virtual library of more than 10,000 compounds is enumerated using an in-house enumeration tool. Compounds are selected via property predictions or project-specific criteria using other tools (e.g., Spotfire). Selected compounds are uploaded to Design Hub via SDfile import; a custom data column is the reactant compound ID. After synthesis and testing, the best compounds are resynthesized for confirmation and further profiling. From this workflow, BI has learned that uploading library compounds to Design Hub supports teamwork and helps avoid duplicates. The merging procedure makes tracking of resynthesis challenging. Support for full-scale enumeration in Design Hub, with reactant handling, is highly anticipated.
Flexibility to plug in additional custom modules is highly valued. Several new smart assistants have been created and rolled out over the past two years and there are more to come. For example, structural alerts have been added. A medicinal chemistry team was formed to comb through internal data and the external literature, potentially problematic substructures were identified and categorized, and a new smart assistant was created to notify designers that a structure contains a potentially problematic substructure.
BI sees the Design Hub of the future as a central tool integrated into the BI digital landscape and an important part of the company’s best-of-breed strategy. Some interfaces are already established. The first D360 connection was recently enabled, and BI is looking forward to an additional, upcoming integration. Connection to the compound management system helps users find and order building blocks for the lab. There is a smart assistant that makes real-time recommendations from an internal data-driven tool; users can explore further via a drill-down menu. An additional connection is planned to send compounds to an automated retrosynthesis tool. Another will send compounds to a CRO synthesis management tool and receive status updates. Design Hub will be integrated with Boehringer-Ingelheim’s ELN.
Accelerate your drug discovery research. D360 and Design Hub integration
Fabian Rauscher, Scientific Informatics Manager (Europe), Certara
Drug discovery research requires the ability to access and understand biological, substance, logistical, and computational data from a wide variety of sources. A major challenge for scientists is the significant time and resources needed to find and access these data quickly and efficiently in order to make critical decisions. D360 is a self-service discovery platform for data access, design, and analysis. Built-in tools include data visualization and structure and sequence analysis. Collaboration and integration facilities include sharable queries, datasets, annotations, and standard desktop tool integration.
Chemaxon and Certara have many customers in common who successfully integrate Chemaxon products with D360. The Chemaxon products concerned are Compound Registration, which acts as a data source for D360; JChem Microservices as additional calculated properties; JChem Cartridge, to provide chemical structure searching within data queries; MarvinSketch, which can be used with D360 for structure input; a customized compound sketcher created for one customer using Marvin JS; JChem for Office in the area of data export from D360; and now Design Hub.
The workflow Fabian presented starts from data retrieval and analytics, as well as compound design in D360 to the design tracking, and prioritization capabilities in Design Hub. Scientists are able to analyze discovery project data to develop SAR using D360’s data access and analysis capabilities; develop scientific hypotheses for improving compound bioprofile using D360; send information and virtual compounds to Design Hub from D360; capture, prioritize, and track hypotheses and candidate compounds through Design Hub; and track the progress of candidate compound synthesis and design goals with Design Hub. Shortening the DMTA cycle time means fitting more cycles into a year; improving the quality within a cycle means fewer cycles are required to reach a given point. The benefit is getting to market faster.
Fabian demonstrated these new facilities. He found compounds from a kinase project by substructure search and displayed the structures and data in a spreadsheet together with some data analysis plots. He added R-group analysis and plotted R1 versus R2 in a matrix. He scored and picked R-group combinations to enumerate, displayed some enumerated structures, and predicted the activity of the virtual compounds with a Free-Wilson model. He sent the virtual compounds with good prediction to Design Hub.
The strategic partnership between Certara and Chemaxon will accelerate drug discovery research through integrated scientific informatic workflows. Adding Design Hub data into the data catalog of D360 for querying and display in datasets alongside real compounds is a logical next step.
Growing the Design Hub system
András Strácz, Product Owner R&D, Chemaxon
Design Hub allows medicinal chemists to:
- analyze project data
- set a hypothesis or goal
- generate compound ideas
- prioritize ideas against resources
- synthesize compounds
- obtain experimental results
- evaluate the hypothesis, and
- share results.
András focused on functions 1, 4, and 6. There is a new unified view for all project compounds, virtual or real, shared or the user’s own, with both predicted and experimental data. A new Extract, Transform, Load (ETL) import system automatically brings in compounds and aggregated assay data, loading from Chemaxon sources as standard. These features are compatible with Certara D360. In Design Hub, multiple references can be used to compare virtual compounds against project front runners. That feature has been revised. New, rich, plugin-based visualizations are available for comparisons. Caching of results has drastically improved performance. Soon, generic graphs will be available for comparisons with reference compounds. Docking integration has been completely rebuilt to take full advantage of the comparison mode. The docking engine has a bring your own license (BYOL) model because Chemaxon do not have, for example, a physics-based product of their own. András showed an animation of the spreadsheet alongside a radar plot and a docking pose.
Registration has been simplified, linking a parent idea to child variants with differing salt forms and stereochemistry. New features include loose matching at “version” level, ignoring stereo and salt, notifications when substance matches are registered, and automatic approval when a single substance matches for a specified time. There is a single sign-on (SSO) based connection to retrosynthetic analysis of compounds (see the chemical.ai entry later in this report). Local and global machine learning models can be used. These can now be automatically distributed to the drug discovery team with the Trainer Engine.
To scale up synthesis resources, pharmaceutical and biotech companies share compounds with CROs. In the past, a project leader or medicinal chemist had to take on the administrative tasks of sending and receiving synthesis information, and to stay up to date. Design Hub separates out data that must not be seen by CROs and the screen formats have space for the CRO to insert replies.
The kanban project management tool has been revised. Options related to hypothesis, design set, assignment, or axis are scoped to projects. To allow users to spot “stuck” compounds easily, time spent in a certain status is now indicated. There have also been improvements in performance and a host of minor improvements.
András concluded with a number of questions for the future. How might Chemaxon support pharmacophore modeling? How might they reduce noise among competing design ideas? How might they help teams with compounds stuck in synthesis? How might they improve the presentation mode of concepts and project status?
Translating data to predictive models
Ákos Tarcsay, Product Manager of Calculators, Chemaxon
The machine learning (ML) life cycle involves collection of data, and experimenting with them (including standardization, training, visualization, and triage), followed by deployment of a model and retraining. Chemaxon have developed a process to help with this from data ingestion through preprocessing, modeling, and review, to prediction. Ákos talked about building the infrastructure (Figure 3) which includes the Java ML library, Statistical Machine Intelligence and Learning Engine (SMILE).
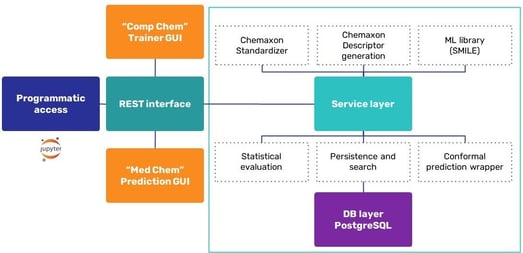
Chemaxon standardization, including handling of salts, solvates, and tautomerism, reportedly outperforms the other approaches.5 Ákos reported very satisfactory results (R2 = 0.9) when he applied the Chemaxon tautomerism algorithm to a small molecule retention time (SMRT) dataset,6 training on 7,000 random, single-tautomer cases, and using a test set of 15,000 compounds (where tautomerization affected 252 cases), in runs with and without tautomerism.
He explored prediction power using activities for a 163 ChEMBL targets from the ChEMBL dataset.7 The dataset was split 10:90 into test and training sets (160,000 total training size, 18,000 total test size) and 30 points from the most recent document year were reserved for the external set. The external set proved particularly challenging. Prediction power for the best models is summarized in Table 1.
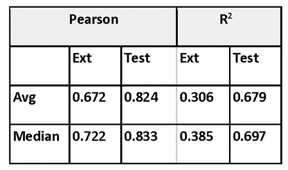
Ákos explored confidence using conformal prediction.8 For a test set of 14,233 out of 17,661, 80.6% of the predictions were within the error bound and for an external set of 3344 out of 4890, 68.4% were within the error bound. In SAMPL7, Chemaxon’s Chemicalize toolkit was tested as an empirical reference method to make macroscopic pKa predictions and it performed better than other methods.
Next, Ákos presented a blood-brain barrier penetration classification use case based on the MoleculeNet dataset.9 A generated random forest model showed high Matthews correlation coefficient (MCC = 0.6) and 0.95 sensitivity using 43 descriptors capturing physicochemical properties and structural descriptors.
Another use case was PAMPA permeability, using the PubChem BioAssay dataset. Ákos standardized and then chose 2029 cases with molecular weight under 800, and with a permeability 100 cut-off using “Phenotype” field. He classified 646 cases as low/medium and 1383 as high. He made a tSNE plot based on MACCS keys and created two clusters. He tried to train on cluster 1 and predict on cluster 2. The base model was a message-passing neural network that resulted in acceptable accuracy (MCC = 0.39) on the training set, but low performance on the test case (MCC = 0.07). The Gradient Tree Boost ensemble model generated with Trainer Engine using 19 descriptors resulted in promising accuracy (MCC = 0.41) on the corresponding test set.
Models can be made available in Design Hub, Chemaxon’s discovery hub connecting chemical series, data, predictions, and chemical project management (Figure 4). Design Hub is service agnostic. You simply set a production flag and then the model is available straightaway to the users.
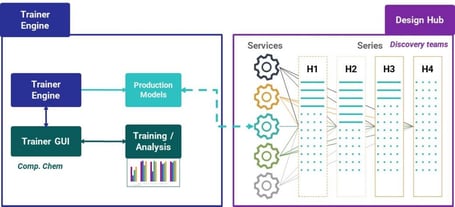
Trainer Engine translates data into reliable models and centralizes model management. Design Hub connects project team members and resources and tracks and manages discovery.
Plan your synthetic routes with ChemAIRS
Ning Xia, CEO, Chemical.AI (Wuhan Zhihua Technology Co., Ltd.)
Organic synthesis is still a rate-limiting factor in drug discovery projects.10 One solution for speeding up the DMTA cycle is computer-aided synthesis planning. AI retrosynthesis tools have already been adopted by some pharmaceutical companies to suggest the shortest and most reliable routes.11,12 There are three different approaches to retrosynthesis: systems based on experts’ rules (examples of which are Chematica, SYNTHIA, and ChemPlanner in SciFindern), purely data-driven tools such as that of Pending-AI and Waller’s lab13 (used in Reaxys), and the ChemAIRS system, developed by Chemical.AI, which does rules extraction and route search by machine learning aided by the knowledge of chemists.
One problem for the purely data-driven approach is that there are few examples of rare reactions from which the system can learn. Another issue is the lack of interpretability. Systems based on rules written by experts are labor-intensive and suffer from the subjective view of the rule writer. ChemAIRS has the advantage of the positive features of both other approaches. Complex synthetic problems are broken down into smaller, simpler ones with chemistry meaning and then data and machine learning are used to solve the problem. The system learns from data automatically and expert knowledge is used to adjust the algorithms. Internal data can be integrated. The results are explainable. The system is easy to debug and enhance. It can be fed with more data to improve the algorithm and develop more products for different scenarios.
Chemical.AI has collaborated with more than seven pharmaceutical companies in the United States and Europe and has more than 10,000 online users. In a C&EN webinar, WuXi AppTech reported an evaluation of nine retrosynthesis software products devising routes for 60 diverse targets. ChemAIRS came out top with a score of 84. The nearest rival scored 64.
Recently, 16 experienced chemists at Shanghai Chemiscal Biotech Co. Ltd. (a subsidiary of Chemical.AI), aided by literature and other tools, challenged ChemAIRS in designing routes to 22 unpublished molecules without chirality. Routes ranged from eight to 14 steps. The competition was judged anonymously by six principal chemists. ChemAIRS took only 8.7 minutes to get multiple routes to a molecule, while chemists took about 1.5 hours per route. In 13 cases out of 22, ChemAIRS’ synthetic feasibility scores were equivalent to or higher than chemists’ scores. ChemAIRS suggested diverse routes: 2-6 different synthesis strategies, including different key steps and intermediates. In one example, a chemist devised a three-step synthesis but ChemAIRS suggested a one-step route. In another case, a chemist’s route had a site selectivity problem but the ChemAIRS route did not.
ChemAIRS is very fast, it can show similar references to the target reactions, it produces a protecting group strategy, it identifies chiral centers and makes the right transformation, and it suggests multiple, diverse strategies. Starting materials and prices are stored. ChemAIRS is easy to use. Structures are entered using Marvin JS. Suggested routes are ordered by feasibility score and coloring distinguishes predicted routes and known compounds.
In one example, ChemAIRS suggested alternative routes, not known by the chemist, for synthesizing the skeleton in a key step and it shortened two other steps into one. ChemAIRS can design routes for difficult targets. For example, chemists failed to find a route to a molecule with a nonaromatic ring and three chiral centers but ChemAIRS suggested 10 routes, two of which were found to be quite feasible. ChemAIRS handles chirality well. In a third example, ChemAIRS found a known literature route to the target but also found two routes of comparable length with different strategies. The first route was more feasible than the literature one and used cheaper starting materials. The second route was two steps shorter than the original one, the steps were feasible, and the starting materials were cheaper. ChemAIRS finds more economic, less risky routes.
A molecule can be sent from Design Hub to ChemAIRS. Once routes are found, the information can be sent back to Design Hub. Chemical.AI has other tools to accelerate preclinical drug research. One is a tool to evaluate the synthetic accessibility of a library of molecules. This has been compared favorably with SAscore14 and Merck meanComplexity.15 In-house data such as building blocks and reactions from ELNs can be integrated in ChemAIRS and more rules can be derived. ChemAIRS can also carry out forward synthesis from a scaffold to produce an enumerated library based on feasible reaction rules and given building blocks. Chemical.AI also offers Mind Map for visualizing a network of routes. It is especially useful for discovering novel reaction routes.
Reaxys predictive retrosynthesis: development, use, and adoption by chemists
Markus Fischer, Product Manager, Elsevier
Data-driven approaches can improve synthesis planning. The Reaxys team and Mark Waller have collaborated to develop predictive retrosynthesis that takes advantage of best-in-class reaction data and ML technologies. Waller’s team wrote one of the most cited works in the field of predictive retrosynthesis,16 reporting a predictive model based on neural networks and Monte Carlo tree search, that learns transformation rules from data, learns to prioritize rules and predict reactions, and uses modern, efficient search methods.
High quality data is essential in such applications, so Elsevier used Chemaxon’s Standardizer for data preparation and canonicalization of chemical structures. Customer centricity is at the heart of design and development of Reaxys predictive retrosynthesis. Pharmaceutical companies defined the most important features of a synthesis planning platform to encourage adoption:
- an easy-to-use and intuitive interface for interaction with the routes
- a scoring system for ranking routes
- termination of routes at purchasable starting materials
- a method to explore the literature precedents associated with route suggestions
- a facility for the user to define the bonds to be broken to guide the search
- identification of functional group incompatibilities and unstable compounds and proposal of protecting group strategies to bypass these complications.
In Reaxys predictive retrosynthesis, researchers simply enter a target molecule in Marvin JS and hit the “synthesize” button. An overview of routes then provides a high-level preview of the published and predicted routes for the target molecule all in one view. Commercially available starting materials are indicated with a shopping basket icon. The number of steps in the route is stated and users are presented with an interactive and graphical route overview to assess complexity.
A tree view of each route provides more details (using Chemaxon’s Marvin editor), including underlying literature precedents and experimental procedures. Users can easily toggle between routes, without the need to navigate between multiple windows. A route can be extended at any step by adding one reaction step at a time (published or predicted). Results can be exported in multiple formats for sharing or adding to ELNs.
Erick Carreira’s lab at the Swiss Federal Institute of Technology in Zurich has evaluated the Reaxys retrosynthesis software. Seven teams were involved, with three or four researchers and at least one senior and one junior researcher in each team. Thirty structurally diverse druglike molecules and natural products were selected as targets. The teams created a synthesis route without predictive retrosynthesis, assessed its scientific viability, and estimated the time and effort required for optimal route generation. They then created a synthesis route using predictive retrosynthesis, compared the predicted route to the expert-generated route, gathered new ideas, and measured the time and effort required.
For one druglike molecule, the retrosynthesis tool proposed an interesting approach and a robust route, with reaction conditions shown. The researchers concluded that human-generated routes tended to be longer as academics tend to focus on catalog chemicals. In industry, chemists might be able to purchase more complex starting materials from CROs.
Another target was a small natural product fragment. The retrosynthesis route was fairly similar to the route devised by a human and it correctly proposed key steps. More work is needed here. For a second natural product, the retrosynthesis route may not have been fully feasible, but the first disconnection proposed was innovative and something that would not have been obvious. The system helps to remove human bias and to explore beyond the obvious reactions. For molecules of such complexity, the ability to custom-build a route by adding one step at a time is a helpful option.
Carreira’s teams concluded that the retrosynthesis tool is user friendly and intuitive for chemists who are familiar with Reaxys. It makes robust predictions for druglike molecules, but chemists do need to review predicted routes and make small adjustments as required. The software saves time by designing synthesis routes and getting literature references and ideas for conditions that can be used. Some suggested steps are very innovative and can be applied in a human-assisted synthesis. Complex molecules such as natural products present a challenge, so full routes might not be provided, but it is possible to get innovative disconnections for some steps. Elsevier plans future enhancements to Reaxys retrosynthesis in collaboration with Chemaxon.
Aiding patent drafting with Markush Editor
Béla Pukánszky, Business Analyst, Chemaxon
Markush claims are important. Between 1994 and 2013, out of 3,704,996 U.S. patents, 468,262 had Markush claims, which often cover more than 1010 structures. Manual patent drafting requires highly skilled professionals. It is tedious, time-consuming, and error-prone. The expert receives a list of compounds, identifies a scaffold, formulates R-groups, and builds up a claim hierarchy. The patent application is finalized by adding a list of examples, adding a detailed description, and completing the formal requirements.
Computer-aided patent drafting can carry out scaffold detection, R-group formulation, interactive editing, search and visualization, and draft generation. Chemaxon’s Markush Editor can automatically generate an initial Markush structure from a list of compounds. Generalizations and modifications can be done in an intuitive editor (Figure 5).
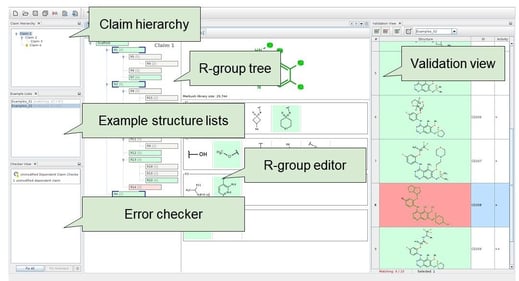
Markush Editor supports various input formats (e.g., SDfile and CSV). It has rich Markush functionality: R-groups, homologies, position variation, repeating units, and multipliers, etc. Visualization is informative and easily understandable. It offers interactive validation against examples (or prior art) and immediate feedback about any inconsistencies or drawing errors. Claims can be exported in English or Japanese. An exemplified structures list is generated with IUPAC names, structure images, and additional information such as assay data. Exported structures are editable in MS Word with JChem for Office. This revolutionary new approach of chemistry patent drafting saves time, avoids mistakes, and checks any structures against your patents easily. The result is better quality patents and competitive advantage in protecting your intellectual property rights.
Markush Editor is being continuously developed. New features include creating dependent claims based on selected examples; customizing exported claim drafts with different journal styles; handling multiple structure lists; exporting additional data to the draft document. Planned new functionalities are improved claim hierarchy creation and generalization workflow; and coverage analysis to help properly support all parts of the claims with examples.
Claim drafting, reverse engineering
Alberto Bertucco, Senior Patent Counsel, Bayer Intellectual Property
Alberto has compared the “manual work” of the patent drafter with the computer-aided alternative by taking a published patent application and partially reengineering the claims with Markush Editor. As an example, he used WO2018134148, a published international patent application filed on January 15, 2018. It claims 2 priorities 17152508.2 EP (January 20, 2017) and 17179210.4 EP (June 30, 2017). With different priority, 199 exemplary compounds are covered. These are spiro- and not-spiro heterocycles under the same general formula. There are several intermediate generalizations: those in claim 1 (the main claim) have general formula A, those in claim 2 (dependent on claim 1) have general formula IA, and those in claim 3 (dependent on claim 1) have general formula B. Alberto fed the 199 compounds, divided in three lists according to the relevant filing date, into Markush Editor.
For general formula A, the software first proposed a general formula with R1 incorporated in the scaffold (Figure 6) and 11 clusters of R2 groups. WO2018134148 had only eight clusters. Markush Editor proposed 25 R-groups (R1-R25), but there were only nine in general formula A (Figure 6) according to the patent. Alberto re-elaborated the proposed general formula in order to align it to general formula A of the patent and he aligned the proposed R-group definitions and numbering with those according to claim 1. He similarly processed formula IA and formula B in Markush Editor, producing claims 2 and 3 as subsidiaries in a hierarchy of claim 1, with general formula A.
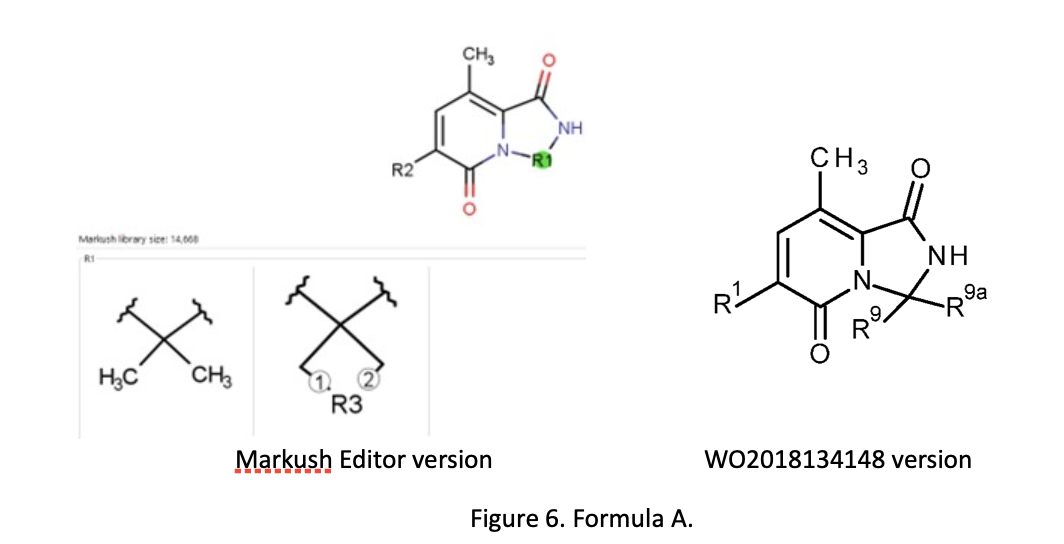
He concluded that Markush Editor manages different compounds lists in a very efficient way making it possible to analyze and keep all compounds under a suitable general formula. The draft of a general formula is facilitated as well as the design of subformulas based on the number of examples. The optimization of the main general formula and the appropriate definition of the R-groups still require some “manual skills”. The claims drafted can be exported in a Word document and optimized for filing.
Compliance Checker and cHemTS
Elke Hofmann and Micha Hofmann, Research & Lab Informatics, Merck KGaA, Darmstadt
Research & Lab Informatics within Merck Healthcare R&D owns and shapes the Merck Healthcare Research Database as the core data and knowledge repository for discovery data and delivers end-to-end scientific IT solutions. Merck has two major internal research hubs and an increasing number of externalized activities involving CROs. One of the many challenges in such a global set up is the compound exchange among internal and external research sites. The effectiveness of compound shipment is crucial and besides the physical parts of the shipment process, legal aspects such as controlled substances and customs law must be considered. There are more than 1,100 shipments per year from one of the major research hubs, in Darmstadt, to more than 30 countries. The use of controlled substances is regulated by governments under internationally agreed schedules and is incorporated into national laws, often with significant expansions. The legislation has an impact on compound inventories at the two major Merck research sites in Darmstadt and Boston.
Merck therefore evaluated several potential software systems that might address the problem. They tested accuracy using example data with a known number of expected hits. They considered whether the countries covered were relevant. The chosen system needed to allow for automatic and interactive processing for both single and multiple compounds, to require minimal effort for installation and running, and to support of a wide range of chemical structure formats. Integration into existing tools had to be as easy as possible. Merck selected Chemaxon’s Compliance Checker.
Compliance Checker supports the healthcare research compound shipment by automated, batch processing of research compounds, and reagent inventories and catalogs. For research compounds, results are propagated to the compound logistics system as “shipment restriction”. Compliance Checker is integrated in registration of research compounds, compound acquisition campaigns, and compound design. Interactive processing uses Marvin and Excel add-ons.
After rollout, Merck found that integration into Marvin for end-user single checks was not as easy as expected: a JSON to HTML wrapper was needed to make the output human-readable, Also, the internal effort for continuous updates of the knowledge base and for continuous checking and rechecking of compound repositories was higher than expected, and system stability was not as good as expected.
As a result, Chemaxon has increased system stability, developed an HTML endpoint for integration into Marvin, runs Compliance Checker as an in-house managed service, and developed a custom application for automated database check which permanently checks new compounds and does a complete recheck after knowledge base updates. Compliance Checker is currently integrated with Merck in-house data and processes for research compounds and reagents, and for acquisition campaigns, where a compound shopping tool avoids buying controlled compounds. All internal compound stores have been checked and all controlled compounds have been discarded.
The Harmonized Tariff Schedule (HTS) is used by over 200 countries, territories, or customs unions around the world to assign a number to each product to establish the appropriate duty that should be paid when goods are imported into a country. Chemaxon’s cHemTS automates the time-consuming process of assigning the correct HTS codes to substances during international shipment.
In 2020, Merck decided to start as early adopters with an “extended evaluation” period. Interestingly, Merck functions from other sectors had an even stronger interest than Healthcare in the topic and played a big part in the evaluation, which focused on features similar to those studied in the Compliance Checker evaluation. In 2021, based on positive feedback from users, Merck decided to add cHemTS to the permanent application portfolio. As regards accuracy, the tested trust level had a result of much greater than 95. Now, processing time has been reduced, quality has increased, and automated conversion of name or CAS RN to structure is used for double checks. A user states, “Our group will be able to perform their work tasks in the required time and quality only with tools like cHemTS”.
Chemaxon runs cHemTS as an in-house managed service for Merck, where it is integrated with in-house data and processes. Shipments of research compounds rely on HTS codes generated by cHemTS and soon, shipment of liquid crystals and OLED compounds will also rely on the system. A cHemTS REST service is used in automation platforms such as KNIME for the building block explorer.
Merck has requested a few improvements to cHemTS. Integration of a larger CAS repository would be helpful. Currently, Chemaxon integrates the CACTUS webservice to resolve CAS RNs into structures but many CAS numbers cannot be converted, so the database is by no means complete. Polymer compounds, ethanol, precious metal compounds, and rare earth elements are mostly in the Chapter 29 tariff, although they belong to other chapters (38, 22, 28). Chemaxon started with Chapter 29, and they are now gradually integrating the other chapters. This is documented but some Merck users overlooked this. HTS codes for compounds which should have tariffs based on their activity, such as vitamins, alkaloids, and hormones, are mostly generated from the chemical structure alone and therefore are often wrong.
Controlled substance detection as a service
Ákos Papp, Product Owner of JChem for Office and Compliance Checker, Chemaxon
The presentation by Elke Hofmann and Micha Hofmann showed how Compliance Checker can be easily integrated when it is deployed on premise, but the same integration is possible if Compliance Checker is hosted in the cloud, and provided as a SaaS solution. SaaS provides many advantages. Eliminating local installation steps reduces upfront cost, and the software can be ready for integration much faster, because all that is needed is minor configurations and addition of users. Ongoing costs such as maintenance are also borne by the vendor, and the system is automatically updated when a new software version, or new content, is available. Since the price of a subscription is not much higher than the cost of a software license, and since for SaaS, Chemaxon rents Amazon Web Services (AWS) resources, provides high availability (HA), and includes maintenance etc., the benefits of SaaS outweigh the extra cost of the subscription.
Chemaxon have several different SaaS offerings, but Ákos spoke only about the Compliance Checker Software as a Service Premium subscription which provides:
- global access with an unlimited number of users
- HA of 99.9% by running multiple, redundant Amazon Web Services
- up to 200 molecules/sec performance on a small molecule druglike compound collection
- cloud hosted data storage
- premium support and accelerated development.
When a regulation change is published, the whole compound repository must be rechecked as soon as possible, but this is usually postponed until the weekend. Chemaxon tested the premium subscription service to see whether rechecking could be completed over the course of a weekend. The Molport database, consisting of about 46 million unique structures, was selected as the dataset. The structures were run in 94 CSV files each comprising half a million SMILES and IDs, against Chemaxon’s full range of regulations, containing 18 country-specific and six international regulations. The work was successfully completed over the course of a weekend. cHemTS is also available by SaaS subscription, either standalone or as a module of Compliance Checker.
In the near future, Compliance Checker will detect compounds substantially similar to directly controlled substances. The U.S. Federal Analogue Act – 21 U.S.C. § 813 also controls compounds that are substantially similar to directly controlled substances. Using standard chemical similarity searching presents a challenge here because, ultimately, a human expert will decide if the compound in question is really similar or not to the controlled one.
The Extended Connectivity Fingerprint (ECFP)17 is one of the most effective fingerprints used as a descriptor and you can find many published articles where a single similarity cutoff value is recommended for a certain activity class. In a study by Franco et al.,18 143 experts were asked to rate 100 pairs of compounds as similar or not, and their scores were compared to the similarity scores generated by ECFP4 fingerprints. Although there are “gray area” examples, the results show that ECFP4 could certainly be a strong starting point for Chemaxon’s similarity method.
In addition, Chemaxon found a few issues with ECFP4 itself. For example, testing showed that ECFP4 was limited in its ability to handle rings: phencyclidine (“angel dust”) and a similar compound with a seven-membered ring in place of cyclohexyl are determined to be identical (Tanimoto coefficient = 1) and macrocycles have too high a similarity to smaller rings. Tanimoto coefficient calculations are also biased toward smaller molecules: 3,4-methylenedioxymethamphetamine (“ecstasy”) and a very similar small molecule with an extra methyl group on the side chain have a Tanimoto coefficient of only 0.538. Chemaxon resolved these issues by introducing a proprietary combination of ECFP4 and another similarity method, and created two similarity categories, which enables the software to distinguish molecules highly similar to regulated ones.
A few optimizations were also needed to improve the search speed with these fingerprints. Omitting large molecules (biologics) enables a greater focus on relevant structures. Only the major tautomers of the example molecules are considered. In addition, stereochemistry is ignored (since the Schedule I and II regulations also control all stereoisomers of the molecule).
Trends and Technology
Trends in life science R&D: some Pistoia Alliance perspectives
John Wise, Pistoia Alliance
The Pistoia Alliance is a non-profit organization with over 200 members, more than 110 of which are companies. Chemaxon has been a member since 2009. The alliance is dedicated to lowering the barriers to innovation in life science and healthcare R&D through precompetitive collaboration. It convenes two large conferences and many other events throughout the year. It runs communities for many “hot topics” such as AI and ML, quantum computing, data governance, lab of the future, and natural language processing (NLP) but fundamental to the Pistoia Alliance are its many projects and expert groups managed under a formal legal framework that allows for agile collaboration in biopharma and life sciences. Two examples of expert groups relevant to the Chemaxon User Group include User Experience for Life Sciences UXLS, and controlled substance compliance and shipping.
In 2020, R&D spending in the pharmaceutical industry totaled nearly $200 billion dollars globally. The pharma market is rapidly changing. Payers are responding to aging populations and chronic diseases. Constrained healthcare budgets are impacting spending on pharmaceuticals. There is increasing pressure to fund drugs for rare diseases. Pharmaceutical companies are responding to increasing R&D costs, falling peak sales per asset, an increasing number of biosimilars entering the market, and delays in patient access following market authorization.
Pharma will have to learn much more about how the human body functions at the molecular level and the pathophysiological changes caused by disease; only then will it be able to develop a better understanding of how to modify or reverse these changes. This is a huge undertaking and one that pharma cannot complete alone. It will require the support of academia, governments, technology vendors, healthcare providers and the regulators, and patients must also play their part in supplying data. FDA figures show an increasing trend toward biologicals. Electronic health information (real-world evidence) can add value. The Nuffield Council on Bioethics has published a report on the collection, linking, and use of data in biomedical research and healthcare.
Many years ago, said Orison Swett Marden said: “No employer today is independent of those about him. He cannot succeed alone, no matter how great his ability or capital. Business today is more than ever a question of cooperation.” The same is true today. The CERN Large Hadron Collider is a recent example of successful collaboration between ostensibly competitive organizations. The Pistoia Alliance is another.
The Pistoia Alliance has published a report entitled “2030 Life Sciences and Health Go Digital”. Hypertension, obesity, noncompliance, diabetes, and asthma etc. (“HONDAs”) are imposing a significant burden on world healthcare resources. The report draws attention to some innovative technologies and patient-centric innovations. In a video, Severin Schwan of Roche notes that society has failed to invest in preventative medicine. There was no testing infrastructure until the recent pandemic struck. The early detection of cancer is particularly important. Other items in the report consider the potential of quantum computing to transform drug discovery and indeed pharma supply chains, the emerging significance of the microbiome, about which the Pistoia Alliance is running a project, and the urgent need to address a widening skills gap not least in data science.
The alliance has three strategic themes: improving the efficiency and effectiveness of R&D (including interconnectivity and data workflows from the lab and ELN to storage of data in FAIR format), emerging sciences and technology (such as quantum computing, AI, ML and NLP, and the microbiome), and empowering the patient (identification of medicinal products (IDMP) ontologies, secondary use of clinical data, and Informed consent with blockchain). Hierarchical Editing Language for Macromolecules (HELM) is a flagship project. Semantic Enrichment of ELN Data (SEED), FAIR implementation, and the Chemical Safety Library are others, among many. Recently, a diversity and inclusion in STEM program has begun.
John presented a list of Pistoia publications, many of them in Drug Discovery Today or Drug Discovery World. He concluded with a movie of the Pistoia Alliance 2018 “Hack the Lab” hackathon highlighting the exciting possibilities of the connected lab and standards for integration of instruments.
Panel discussion
Chair: John Wise (JW). Panelists: Thomas Balkinas of Amazon (TB); Richard Jones of Chemaxon (RJ); Wendy Warr of Wendy Warr & Associates (WW)
JW: Richard, does the cloud offer an opportunity or a threat to a cheminformatics company? Is it a risk to push your company to the cloud and how do you align your company around it?
RJ: It's more of a risk not to focus on the cloud. You just have to look at the world around you to see the positive effect the cloud is having. Within our own industry we see companies changing from “cloud last” to “cloud first” but you have to do this right. You can’t go for a lift and shift approach of your software: you have to look at the capabilities and innovations you can generate through the cloud for your customers and focus that way.
Regarding alignment of the company, we follow the same approach our customers have, namely, focus first on the business case and use this to get the leadership on board. You then look for funding and people to help you realize it. The biggest challenge will always be the cultural shift that this brings. We are in the lucky position where we generate a lot of revenue from legacy products, but this can lead you to become stagnant. We need to move with the times and innovate and this requires education and reorientation throughout the entire company.
JW: Should users see the cloud as the way forward? Does the journey to the cloud have any major challenges? Does the cloud (versus on-premise solutions) help with collaboration and cybersecurity?
TB: It is very important to migrate user needs, not just products, to the cloud. We at AWS want our customers to choose us because we offer the broadest and deepest set of services, best partner solutions, and the most secure place to run their business, and at optimal speed and cost.
Question from the floor: What about the fear of vendor lock-ins?
TB: We’ve built our cloud infrastructure on open standards. This means that you have the freedom to move your own data wherever you want. In fact, the same tools we offer to migrate into the AWS cloud can be used to help you migrate out of it. We’re not in the business of restricting how customers use their technology. This reduces the sense of risk, and ultimately makes for a healthier and more productive partnership. The cloud should be a catalyst for innovation, not another name for penalizing lock-ins. There is a blog post with a full analysis of AWS’s position on vendor lock-in.
JW: What role can AI play in today’s cheminformatics?
WW: Not many years ago, Johnny Gasteiger (who in 1999 wrote a seminal book19 about neural networks) said that AI was a great way of getting grants but was not much use in practice. Things are slowly changing. There has been a lot of interesting work recently on the use of AI and ML in retrosynthesis and other reaction informatics fields but is still difficult to persuade medicinal chemists of the value of these new technologies. Another innovation in drug discovery is the exploration of chemical space and the use of ultralarge compound collections.20 GSK, for example, has constructed a chemical space of 1026 virtual compounds (not with fully enumerated structures, though).
JW: Other innovations include technology platforms to support the new research modalities (oligos, antibody-drug conjugates, and cell and gene therapies), and in my talk earlier today, I emphasized the importance of collaboration in support of innovation. Is there a use of blockchain technology in biopharma?
WW: Actually, there is one significant collaborative blockchain venture: it’s called Machine Learning Ledger Orchestration for Drug Discovery (MELLODDY). Ten pharmas are involved in federated, privacy-preserving machine learning. Each partner has a private AWS cloud.
TB: Blockchain is also being used for managing the threat of counterfeit drugs, in particular where supply chains go through difficult countries, and it can prevent drug waste due to lack of trust in the provenance of the supply chain.
Question from the floor: Is it easier to train data scientists to be chemists or to train chemists to be data scientists?
WW: Major university libraries now have data librarians. And at the recent AI4SD meeting in Southampton there was discussion on teaching chemistry undergraduates about Python.
RJ: We should be starting the education in high school. In some schools in England, they are already giving pupils Raspberry Pis to program which is a great idea. It does open up issues with equality and access though, but perhaps governments can get involved.
JW: A final thought: sustainability is an issue, including green chemistry, which leads us to the concept of “green cloud”. The cloud needs to be green too!
Cheminformatics and bioinformatics in the cloud
Gábor Pécsy, CTO, Chemaxon
In early 2021, Synergy Research Group published some analysis of the megatrends of the previous decade which revealed that during that period, enterprise data center spending stagnated while cloud infrastructure spending grew superlinearly, exceeding the former by the end of the 2020. Before the pandemic, Gartner forecast that worldwide end-user spending on public cloud services would grow 20.4% in 2022 to total $494.7 billion. The need for remote collaboration and support for home office gave a huge boost to cloud services and Gartner has now increased the forecast figure to nearly $500 billion.
Chemaxon is ready to serve customers’ needs in the cloud. By 2025, the company wants to renew its portfolio and provide cloud-based solutions to all its clients. There are many reasons for the success of cloud, and numerous benefits that drive this migration. COVID showed us the importance of tools that enable collaboration in a geographically distributed team. Managing the cost of operation and the ability to grow the system gradually as needs grow help to run businesses predictably. The ability to rent people’s skills or computational resources for the period of need alone makes it possible to solve problems that would be prohibitively expensive or impractical in a private setup, and last, but not least, the SaaS model supports innovation and agile software development methods best and hence it enables the fastest value creation for customers.
Cloud computing is a paradigm, a distinct set of concepts or thought patterns, including theories, research methods, postulates, and standards for what constitute legitimate contributions to a field. This means that to reap all the benefits of cloud computing, Chemaxon have to build cloud-native applications using cloud-compatible methods. The company needs to adapt to those changes, and learn new concepts, pattern, theories, and standards.
First and foremost, software-as-a-service shifts responsibilities. In the past, Chemaxon delivered components that were run by customers who bore a big part of the responsibility for the security of their data. In the SaaS world, customers trust Chemaxon with that responsibility. Chemaxon take this responsibility very seriously. They chose AWS, the most mature cloud provider. They trained their developers and product managers on secure software development and security testing. They introduced a DevSecOps culture: security concerns are tackled from the very early phases of the software lifecycle. They constantly monitor and analyze their code base for vulnerabilities and they fix any they find. In the past two years, the company has built an information security management system compliant with the ISO 27001 standard. Chemaxon continuously strengthen their security system and improve processes. In particular, they invest in automating processes to minimize the chance of human mistakes. They plan to get further certifications in future.
To ensure high quality and smooth operations, the company introduced a DevOps culture into the development teams. This means that the development team took responsibility for the operation of their systems. This enables them to correct production issues more efficiently and it also gives them the insights required to tune their systems to customers’ needs. Thus, more customer value can be delivered in a shorter time and with higher quality. Applications not built for the cloud will fail to deliver the values of the cloud. You should not lift and shift non-cloud applications. Migration to the cloud means migration of user needs not of actual products.
Chemaxon chose AWS as cloud provider because AWS offer a rich set of services which greatly simplify the development of cloud-native services. AWS understand cloud and know how best to support developers. They offer alternatives to enable developers to pick the best solution for their particular use case.
Chemaxon’s current cartridge solutions are the best available technology for relational databases, but AWS offer at least half a dozen different storage technologies tailored for different use cases. When solving a molecule search problem in AWS, we should not think in terms of setting up the database server in the cloud with the cartridge. While it can be a valid solution, it may not be the optimal one, in terms of operation or scalability. We should pick the right storage technology instead and provide the required chemistry capabilities to achieve a much higher quality service. Chemaxon are building their new platform product along these principles to ensure that they deliver the best cloud experience for customers. Some experiments or proofs of concept are shown in Figure 7. The platform will then add the required chemistry and biology capabilities to the cloud and using them Chemaxon will deliver cloud-native solutions to customers’ problems that can provide the full cloud experience.
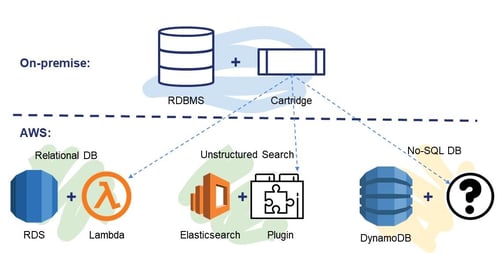
In short, cloud is here, it is inevitable, and it offers very important benefits for everyone. It is a paradigm: you have to adapt your methods and technologies. Migration to cloud means migrating user needs and offering cloud-native solutions. Chemaxon is committed and ready to embrace the cloud and meet the challenges of cloud migration.
AWS for Life Sciences
Thomas Balkizas, EMEA Head of HCLS Partner Solutions, Amazon Web Services
For almost a decade, AWS has helped global life sciences companies migrate to and thrive in the cloud. Once the cloud infrastructure is put in place, innovation can move at a much faster pace. A prime example of business value can be found in Deloitte’s January 2022 report which reported a significant increase in investment levels, related to digitization of labs and the ability of organizations to move their clinical trials to remote and decentralized capabilities. Deloitte credits the cloud for enabling this technology shift.
The strategic journey to build a secure, compliant, scalable, and business-driven infrastructure starts with migration to the cloud. For example, Takeda announced plans in 2020 to move the organization’s applications to the cloud through Accenture, an AWS partner. Takeda was able to move 80% of its applications to AWS in just 18 months, removing non-differentiating technology, reducing the internal data center footprint, and decreasing capital expenditures. Moderna was the first ”born in the cloud” biopharma and deployed the first GxP compliant process built on AWS. Eli Lilly unified data silos to enable more informed decision making, accelerate research, and foster stronger internal collaborations. After data and applications have migrated, and been secured and unified, true innovation can begin.
Thomas cited some common reasons for choosing AWS:
- highest performant ML and high-performance computing (HPC) capabilities, and extensive global infrastructure
- support of more security standards and compliance certifications than any other offering
- wide breadth of cost- saving tools, including automated cost-savings programs for storage
- ease of finding the best partners, tools, and resources with AWS for Life Sciences.
Some organizations want to customize everything while others prefer to buy out of the box. AWS technology partners such as Philips and TetraSciences offer out-of-the-box solutions. Validated consulting partners such as BioTeam and Deloitte specialize in implementing life sciences workloads on AWS. AWS for Life Sciences-is a curated portfolio of AWS and AWS partner solutions and tools purpose-built for the life sciences industry, supporting innovation and operational excellence across the value chain.
R&D applications include automation of repetitive tasks; ensuring GxP compliance and ensuring data are handled, stored, and documented correctly; connectivity with third parties and between instruments and the cloud; and ability to implement new technologies such as next-generation sequencing and Cryo-EM. For example, using Amazon SageMaker, Janssen implemented an automated ML operations process that improved the accuracy of model predictions by 21% and increased the speed of feature engineering by approximately 700%. Using HPC, Relay Therapeutics is able to perform the analysis of billions of compounds in one day, solving the CPU cost challenge with the elastic capacity of Amazon Elastic Compute Cloud (Amazon EC2) Spot Instances. ThermoFisher uses its AWS IoT Core infrastructure to power ThermoFisher Connect, a computing platform benefiting from the power of the cloud, connected devices, and advanced analytics. Biogen uses AWS’ secure global infrastructure to enable secure collaboration with its partners around the globe to analyze data from the UK Biobank.
Further down the value chain are clinical trials. The ability to conduct clinical trials remotely dramatically improves enrollment success and ensures greater diversity and equity in trials. AWS have also been working with customers to enhance trial protocol development to avoid the costly need for amendments. Additionally, AWS are helping customers to integrate wearable device data securely and collaborate with CROs and other third parties. Finally, AWS are helping customers to automate and optimize their regulatory submission processes.
Bristol-Myers Squibb was able to run advanced clinical trial simulations using AWS HPC, reducing their total analysis time by 98%, and reducing the number of subjects needed for the trial by one third. Using AWS IoT capabilities, Evidation has created an infrastructure to analyze passively collected health data from hundreds of thousands of sensors in devices such as smartphones. TrialSpark has used a wide range of AWS capabilities, including EC2 for accelerating trial recruitment processes. Using Amazon Comprehend Medical, Fred Hutch microbiome researchers were able more quickly to identify patients for clinical trials who might benefit from specific cancer therapies.
In manufacturing and supply chain, AWS helps customers by unifying data silos and identifying opportunities for optimization, cost savings, and working capital improvements. The data are also useful for enhancing forecasting. Additionally, instrument connectivity and access to scalable HPC and ML workloads enables organizations to implement new processes for large molecule drug manufacturing. Novartis have created sophisticated ML models to advance operational forecasting for manufacturing and supply chain. Novo Nordisk works with Aizon, an AWS for Life Sciences partner, to put IoT sensors on the shop floor. Merck used AWS to build the Change Assessment Knowledge Engine which is powered by an Amazon Neptune graph database containing information about Merck’s supply chain and regulatory operating environment. Using SAP S/4HANA in a GxP-validated environment in AWS, Moderna built a GMP-compliant manufacturing facility that delivers speed, scalability, and lower costs.
AWS also has customers in commercial and medical affairs. AstraZeneca’s data scientists in the commercial space use AWS to accelerate insights, using Amazon SageMaker and cataloging services. Propeller Health, an AWS customer, was one of the first companies to insert sensors into asthma inhalers. Takeda used advanced analytics to collect and analyze real-world evidence concerning nonalcoholic steatohepatitis. In the area of pharmacovigilance, AWS have customers who have used Amazon Comprehend Medical or Amazon SageMaker to analyze adverse events. Eli Lilly used Comprehend Medical to reduce down to seconds the time it took to analyze an event.
AWS supports 96 security standards and compliance certifications. Bristol Myers Squibb runs SAP on the cloud, using AWS CloudFormation to create a consistent, scalable, and repeatable compliance process. Idorsia uses AWS Lambda to ensure GxP compliance by executing regulated tasks. Moderna have demonstrated the importance of creating a secure and compliant framework that can scale as needed.
AWS has published an e-book on the web entitled The Life Sciences Guide to Cloud Modernization outlining why leading life sciences organizations are turning to the cloud and describing a cloud migration strategy.
JChem Choral cartridge migration use case
Lili Herendi, Business Analyst, Chemaxon and Róbert Wagner, Software Developer, Chemaxon
Róbert described the technical aspects of a use case where migration from the BIOVIA cartridge to the JChem Choral cartridge was carried out. With the JChem Oracle Cartridge (JOC) reaction mapping was costly, especially for biological structures, but this was mostly during indexing. Unfortunately, for certain structures during search time, non-persistable structures were produced. The team wanted first to establish which cartridge was the better for this use case, JChem Oracle Cartridge (JOC) or Choral, the newer version of Chemaxon’s cartridge.
In the cartridges, an Oracle-specific persistence problem can appear due to the way preprocessed molecules are stored. Due to size limitations in JOC, storing molecules which are large or have certain specific features does not allow preprocessing to take place. If there are too many of these molecules present at the same time, the preprocessing stage will take place again, which eventually results in extending the search time. On the other hand, Choral does not have a set size limit, which overcomes this issue more easily.
The JChem Oracle Cartridge (JOC) had some advantages: Chemaxon had more experience with migration and the “halt-on-error” issue was already resolved. “Halt-on error” needs to be set in order for the indexing to continue and not to stop if there is an erroneous structure or problem with the molecules. This is a flag in JOC which Chemaxon implemented, and the user can manually set it. Choral works as if “halt-on-error” is already set. Almost all users set the halt to “on” so Chemaxon thought that this could be a default setting.
Choral had more advantages. There was no reaction mapping of target structures during search time (thus no null cd_smiles); reaction property removal during standardization was built-in so fewer custom configurations were needed; and the solution was future-proof. On the other hand, development was needed to fix the halt-on-error flagging.
The user site migration test environment involved access to the Oracle database and test server for 705,000 molecules and 826,000 reactions. Basic aromatization was used in standardization during installation and setup. The halt-on-error flag was switched appropriately and four molecules (with many polymer component brackets) were not indexed.
Lili related a comparison of the JChem Choral and BIOVIA cartridges (Table 2). The test cases were provided by the client in a form of query molecules and search SQLs with a Symyx syntax and Symyx results. Chemaxon wrote an automatic testing process based on the Symyx syntax with the additional coding required. One of the 154 test cases gave a false negative hit in a polymer reaction, a problem resolved in the latest version of Choral. Thus, Table 2 indicates 153 test cases in all.
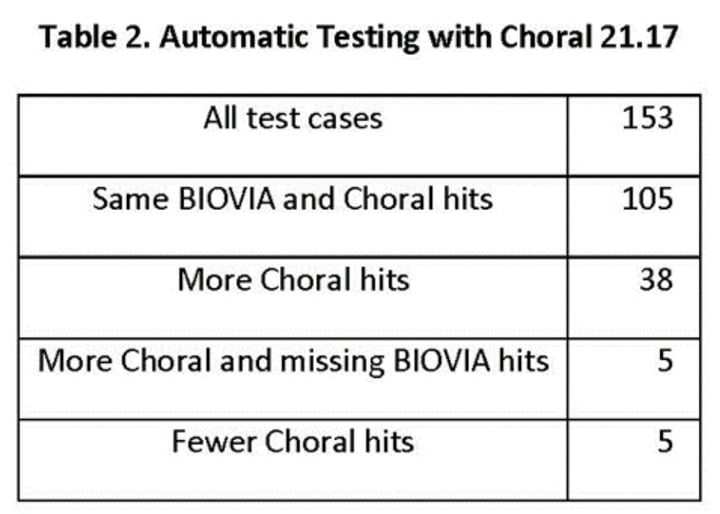
The test results were produced on different states of the database: the expected test results and the test database were different and not fully in sync. Multiple mapping is not supported with reaction properties in Choral so in some query reactions the reaction property was not correct in the reaction (Figure 8).
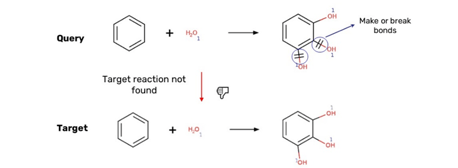
There was an “any atom” and general aromatization issue (Figure 9). The solution might be to just set the aromatization to basic in the cartridge.
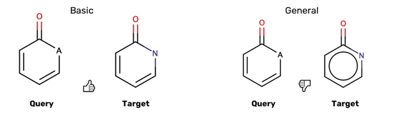
Out of 154 test cases, 144 presented no problem. Chemaxon analyzed the 10 cases with mismatching (fewer) Choral hits. Six were caused by the test data mismatch problem. Three were related to fully mapped, but stoichiometrically incorrect reactions (the multiple mapping problem). One was a case where a single bond in the query did not match to the aromatic bond in the target (Figure 10).
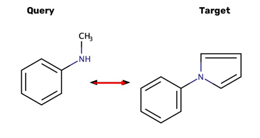
Chemaxon Object Notation
István Őri, Technical Lead, Chemaxon
Among the various file formats, XML is well structured, can be parsed with common tools, and is a self-contained specification, but it has XML external entity injection (XXE) vulnerability and other security issues, and there are size issues. Binary formats store a lot of data in a compact size, they can be very fast and each one works very well in its native environment, but each one causes problems outside of its native environment, it is hard to extract data from binaries, and it is hard to write data into binaries. FORTRAN “serialized” formats such as CTfile are surprisingly easy to work with and incredibly popular but a special tool is needed to read and write them, and they lack some chemical and document features. Encoded formats such as SMILES and InChI are compact, but they too require special tools and have missing features. When used for HTTP requests, XML has security flaws, and many WAFs block it, binary is hard to read and write, and very sensitive to encoding, CTfile formats are sensitive to whitespaces, and encoded formats have some features missing. Outside the field of cheminformatics, JSON beats XML.
CXON (sɪksɒn) is Chemaxon’s new, JSON-based chemical file format. It is suitable for HTTP communication, designed for chemical documents, and easy to integrate. It can be serialized and deserialized with native tools in various programming languages and it is self-documented and rich in features. In future, Chemaxon will offer support for more languages. Java compilation units make it possible to serialize and deserialize CXON files from and into a typed data structure and the same is true of Python. The primary aim is to make CXON as rich in features as MRV and then go beyond that. In addition, Chemaxon will optimize performance in its tools and introduce options such as “pretty print”. Beware, though, that CXON is still in the beta phase, which means that it is possible that backward compatibility may be broken in some cases. Marvin Pro already uses CXON as its primary file format.
Hands-on Workshops
On the first day, there were two short hands-on workshops. In the first, users were able to try out the Marvin Pro chemical editor, the newest generation of the Marvin product family. In the second workshop, users tried out Trainer Engine, the newest addition to Chemaxon Calculators, enabling machine learning techniques to predict data. The meeting ended with a complex laboratory workflow hand-on session where attendees started with a task assignment in Design Hub, created experiments to synthesize the assigned compounds in the new notebook, worked in LabCup’s inventory system (integrated in the notebook), performed a two-step registration in Compound Registration (fixing any errors that occurred), worked with assay results and uploaded them to Assay, ran visualization in Tableau, and returned to Design Hub to analyze the results further.
Partner Session
Gergely Makara of ChemPass spoke about AI-assisted lead discovery in synthetically enabled chemical space. The company has an end-to-end platform with design space components, including SynSpace,21 and analogue cloud analysis, followed by multiparameter optimization (MPO) scoring, ranking, and selection. ChemPass has collaborated with Chemaxon since 2016 and several Chemaxon components such as Reactor, Standardizer, Structure Checker, property Calculators and JChem are built into SynSpace.
Lorena Zara of Discngine spoke about Chemaxon and 3decision, an SaaS platform for structural analytics and knowledge management. It is a protein structure repository of structures, sequences, and associated data, with a collaborative web-based interface that allows users to centralize, visualize, analyze, and annotate protein and ligand structural data. Discngine uses Marvin JS, JChem Web Services and Choral. The company is now moving into biologics enabled by Chemaxon’s Biomolecule Toolkit and BioEddie.
Gábor Radics of LabCup spoke in the partner session but also gave a longer talk summarized earlier in this report. Mcule, represented by Bence Barna, builds three chemical marketplaces for drug discovery: the Mcule database of supplier catalogs, the ULTIMATE virtual universe of 180 million of synthetically accessible compounds, and SYNTHAGORA, a custom synthesis auction site. You can integrate the Mcule database into your in-house system. The Express version of ULTIMATE offers 58 million enumerated, make on demand compounds. Mcule uses Chemaxon’s property Calculators, Compliance Checker, and Marvin JS.
Bérénice Wulbrecht of ONTOFORCE talked about the DISQOVER knowledge platform for life sciences which is integrated with Chemaxon’s Marvin JS, Calculators, and JChem. DISQOVER links any type of data to deliver actionable insights. DISQOVER solves complex use cases over a multitude of heterogeneous data sources with simple, intuitive, and customizable dashboards. Discovering unexpected new insights is a core objective of DISQOVER. At any time, the knowledge graph allows you to follow links to additional, related information for the result set from a search.
Conclusion
The take-home message was without a doubt “cloud, cloud, cloud”. As Richard Jones said, “Chemaxon has a best-in-class, single research platform in the cloud for end-to-end, early-phase drug discovery.” Other big themes were data, and AI and ML, which need high quality, standardized data. In this year’s report, I have put much more emphasis on Chemaxon talks than I usually do, and for good reason. Many new solutions are being launched: an ELN, Marvin Pro, CXON, Trainer Engine in Calculators, and synthesis route design and prediction through partnership with Chemical.AI. In addition, Design Hub has been significantly enhanced. The company’s new culture and strategies were revealed. The addition of a panel discussion to the meeting was an interesting innovation. As usual, there was also plenty of time for networking and meeting up with colleagues again after a two-year hiatus. It was a real pleasure to be back in Budapest. I hope that readers will find value in my summary of this interesting meeting.
References
(1) Vargason, A. M.; Anselmo, A. C.; Mitragotri, S. The evolution of commercial drug delivery technologies. Nat. Biomed. Eng. 2021, 5 (9), 951-967.
(2) Jayatunga, M. K. P.; Xie, W.; Ruder, L.; Schulze, U.; Meier, C. AI in small-molecule drug discovery: a coming wave? Nat. Rev. Drug Discovery 2022, 21 (3), 175-176.
(3) Kuzmina, O.; Hartrick, E.; Marchant, A.; Edwards, E.; Brandt, J. R.; Hoyle, S. Chemical management: storage and inventory in research laboratories. ACS Chem. Health Saf. 2022, 29 (1), 62-71.
(4) Chesnokov, G. A.; Gademann, K. Concise total synthesis of peyssonnoside A. J. Am. Chem. Soc. 2021, 143 (35), 14083-14088.
(5) Dolciami, D.; Villasclaras-Fernandez, E.; Kannas, C.; Meniconi, M.; Al-Lazikani, B.; Antolin, A. A. canSAR chemistry registration and standardization pipeline. J. Cheminf. 2022, 14 (1), 28.
(6) Domingo-Almenara, X.; Guijas, C.; Billings, E.; Montenegro-Burke, J. R.; Uritboonthai, W.; Aisporna, A. E.; Chen, E.; Benton, H. P.; Siuzdak, G. The METLIN small molecule dataset for machine learning-based retention time prediction. Nat. Commun. 2019, 10, 5811.
(7) Lenselink, E. B.; ten Dijke, N.; Bongers, B.; Papadatos, G.; van Vlijmen, H. W. T.; Kowalczyk, W.; Ijzerman, A. P.; van Westen, G. J. P. Beyond the hype: deep neural networks outperform established methods using a ChEMBL bioactivity benchmark set. J. Cheminf. 2017, 9, 45.
(8) Norinder, U.; Carlsson, L.; Boyer, S.; Eklund, M. Introducing conformal prediction in predictive modeling. A transparent and flexible alternative to applicability domain determination. J. Chem. Inf. Model. 2014, 54 (6), 1596-1603.
(9) Wu, Z.; Ramsundar, B.; Feinberg, E. N.; Gomes, J.; Geniesse, C.; Pappu, A. S.; Leswing, K.; Pande, V. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 2018, 9 (2), 513-530.
(10) Blakemore, D. C.; Castro, L.; Churcher, I.; Rees, D. C.; Thomas, A. W.; Wilson, D. M.; Wood, A. Organic synthesis provides opportunities to transform drug discovery. Nat. Chem. 2018, 10 (4), 383-394.
(11) Campos, K. R.; Coleman, P. J.; Alvarez, J. C.; Dreher, S. D.; Garbaccio, R. M.; Terrett, N. K.; Tillyer, R. D.; Truppo, M. D.; Parmee, E. R. The importance of synthetic chemistry in the pharmaceutical industry. Science 2019, 363 (6424), eaat0805.
(12) Struble, T. J.; Alvarez, J. C.; Brown, S. P.; Chytil, M.; Cisar, J.; DesJarlais, R. L.; Engkvist, O.; Frank, S. A.; Greve, D. R.; Griffin, D. J.; Hou, X.; Johannes, J. W.; Kreatsoulas, C.; Lahue, B.; Mathea, M.; Mogk, G.; Nicolaou, C. A.; Palmer, A. D.; Price, D. J.; Robinson, R. I.; Salentin, S.; Xing, L.; Jaakkola, T.; Green, W. H.; Barzilay, R.; Coley, C. W.; Jensen, K. F. Current and future roles of artificial intelligence in medicinal chemistry synthesis. J. Med. Chem. 2020, 63 (16), 8667-8682.
(13) Segler, M. H. S.; Kogej, T.; Tyrchan, C.; Waller, M. P. Generating focused molecule libraries for drug discovery with recurrent neural networks. ACS Cent. Sci. 2018, 4 (1), 120-131.
(14) Ertl, P.; Schuffenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J. Cheminf. 2009, 1, 8.
(15) Sheridan, R. P.; Zorn, N.; Sherer, E. C.; Campeau, L.-C.; Chang, C.; Cumming, J.; Maddess, M. L.; Nantermet, P. G.; Sinz, C. J.; O'Shea, P. D. Modeling a crowdsourced definition of molecular complexity. J. Chem. Inf. Model. 2014, 54 (6), 1604-1616.
(16) Segler, M. H. S.; Preuss, M.; Waller, M. P. Planning chemical syntheses with deep neural networks and symbolic AI. Nature 2018, 555 (7698), 604-610.
(17) Rogers, D.; Hahn, M. Extended-Connectivity Fingerprints. J. Chem. Inf. Model. 2010, 50 (5), 742-754.
(18) Franco, P.; Porta, N.; Holliday, J. D.; Willett, P. The use of 2D fingerprint methods to support the assessment of structural similarity in orphan drug legislation. J. Cheminf. 2014, 6, 5.
(19) Zupan, J.; Gasteiger, J. Neural networks in chemistry and drug design, Wiley-VCH: Weinheim, Germany, 1999.
(20) Warr, W. A.; Nicklaus, M. C.; Nicolaou, C. A.; Rarey, M. Exploration of ultralarge compound collections for drug discovery. J. Chem. Inf. Model. 2022, 62 (9), 2021-2034.
(21) Makara, G. M.; Kovacs, L.; Szabo, I.; Pocze, G. Derivatization design of synthetically accessible space for optimization: in silico synthesis vs deep generative design. ACS Med. Chem. Lett. 2021, 12 (2), 185-194.